BIO 442 MENU
syllabus 
1 - genome
2 - mutate
3 -cell cycle
4 - karyotype
5 - chromoabn
6 -sex-determ
7 -prenatal
8 - mendelian
9 - complex
10 - non-trad
11 - clinical
12 - newborn
13 - teratog 
14 - linkage
15 - DNA prof 
16 - quanti 
17 - links
18 - quizzes
(full title of lecture appears in status bar on the top or at the bottom of your window)
Biology 442 - Human Genetics
Chromosome Analysis: Karyotyping, Banding and FISH
Chromatin refers to "colored" material in the nucleus. We can think of it as chromosomal material. Heterochromatin refers to non transcriptionally active chromatin. There is facultative heterochromatin such as the X chromosome which is capable of being active transcriptionally although it is not always active and constitutive heterochromatin such as the q arm of the Y chromosome or the centromeric heterochromatin which is never transcriptionally active. Euchromatin refers to transcriptionally active chromatin.
The normal or euploid number of chromosomes in human somatic cells is 46. Cells with 46 chromosomes are said to be diploid and the abbreviation used is 2n. There are 22 pairs of autosomes and one pair of sex chromosomes, the X or Y. Females have two X chromosomes and males have an X and a Y. Gametes contain 23 chromosomes, half the number of chromosomes found in somatic cells. They are said to be haploid which is abbreviated n. Fertilization is the fusion of two gametes, a sperm (n) and an egg (n), to form the zygote which has the diploid number (2n). Each parent contributes one member of each homologous pair of chromosomes to the offspring. The egg carries 22 autosomes and an X chromosome while the sperm has 22 autosomes and either an X or a Y. Occasionally, there are tetraploid cells (4n) in the liver. Cancer cells often have an abnormal number of chromosomes (heteroploid) and/or structurally abnormal chromosomes.
Karyotyping
Karyotyping is the process of preparing chromosomes for analysis. The tissues used vary but the most common tissues sampled are lymphocytes (white blood cells), skin fibroblasts, amniotic fluid cells, gonadal tissue, chorionic villi (CVS) and bone marrow. Direct preparations without the use of a mitogen are made on bone marrow, lymphocytes, gonadal tissue, CVS, and tumors. When cultured, cells are placed in a nutrient solution containing amino acids and other nutrients, a pH indicator, antibiotics, with or without mitogens (phytohemaglutinin,PHA, or pokeweed mitogen) and then grown at 37°C in CO2 atmosphere. Lymphocytes are cultured three days and amniotic cells, fibroblasts, CVS, and other fetal tissues are cultured ten days. Colcemid, which inhibits the assembly of microtubules of the spindle, is added prior to harvest to stop mitosis and to collect sufficient cells with prophase-prometaphase chromosomes. Cells may be grown on microscope slides in sterile chambers or they may be attached to the flask in which they are grown. If release of cells from substrate is necessary, trypsin and EDTA are added. The cells are then centrifuged to remove the culture medium and/or the trypsin and EDTA. A hypotonic solution is added to swell the cells followed by centrifugation and fixation in methanol:acetic acid. The cells are dropped onto slides if they were not grown on slides. Most banding procedures require pretreatment before staining. Trypsin and Giemsa produce G-banded chromosomes (GTG).
The slides are scanned for metaphase spreads and usually 10 to 30 cells are analyzed under the microscope by a cytogeneticist. When a good spread (minimum number of overlapping chromosomes) is found, a photograph is taken or the analysis is done by a computer. The chromosomes are arranged in a standard presentation format of longest to shortest. Actually chromosome 21 is smaller than chromosome 22, however, since Trisomy 21 (Down Syndrome) had already been named, it was decided to leave the numbering system as it was. In a standard karyotype, the chromosomes are arranged by size and location of the centromere. There are 7 groups: A, 1 - 3 long metacentrics; B, 4 - 5 long sub-metacentrics; C, 6 - 12 sub-metacentrics; D, 13 - 15 acrocentrics which may have satellites (normal chromosome polymorphisms) on the p arm; E, 16 - 18, sub-metacentrics; F, 19 - 20, short metacentrics; G, 21 - 22, acrocentrics with or without satellites (normal polymorphism) on the p arm; the X chromosome is a sub-metacentric which belongs in the C group and the Y chromosome is a sub-metacentric which can be placed in the G group on the basis of size. The shorter arm is placed on top in a karyotype and is called the p arm, the longer arm is on the bottom and is called the q arm.
1. Metacentric 2. Sub metacentric 3. Acrocentric
There is a standardized way of denoting the bands on banded chromosomes. The karyotypes of normal individuals are almost identical with the exception of some chromosome polymorphisms such as the satellites of the D and G group chromosomes and some heterochromatic regions around the centromeres of 1, 9, 16 and Yq. Each chromosome arm is numbered beginning with the centromere: 11 or 11.1 at the centromere, e.g., 21.2 : Region = 2, Band = 1 and sub band = 0.2
The chromosomes of the great apes are remarkably similar to ours except for some chromosome rearrangements. Chimps, orangutans and gorillas have 48 instead of 46. The human chromosome #2 is the result of a translocation of two chromosomes found in the great apes. There has been a Down Syndrome chimpanzee with an extra small chromosome homologous to our 21.
Human Chromosomes 1, 2, 3, 4 compared to the chimp chromosomes Human Chimpanzee Orangutan Gorilla

Orangutan Whole Chromosome Painting (wpc)
Banding techniques
GTG (pretreatment with trypsin then Giemsa stain) or GTW (Geimsa-trypsin-Wrights); Q (quinicrine, fluorescent bands); R (reverse banding) requires heating in a saline buffer prior to staining with Giemsa or Quinicrine and is often called T (telomeric) banding since it also highlights the telomeres; C (centromere or constitutive heterochromatin) stains the heterochromatin in the centromeres, especially of chromosomes 1, 9, and 16; NOR (nucleolar organizing regions) silver nitrate stains selectively the satellite stalks of the acrocentric chromosomes; sister chromatid exchange (SCE) requires replication in 5 BUdR, an analog of thymidine followed by Höescht stain. SCE is used in the diagnosis of Bloom Syndrome, a genetic disorder that results in chromosome breakage. It is also used as a test for mutagenicity.

GTG banded human chromosomes with banded cartoon along side
C Banding (Centromere or Constitutive Heterochromatin)
Sister Chromatid Exchange (left is normal; right is Bloom Syndrome)
The major dark and light areas starting from the centromere have numbers. Each band is numbered beginning with 11 or 11.1 at the centromere. When you see numbers assigned, the first number is the region, the second is the band within the region, and the third number is a sub-band within the band. For example: 31.2 means 3 is the region, 1 is the band, 0.2 is the sub-band. Regular banding produces about 550 light and dark bands. High resolution banding (HRB) to look for microdeletions requires the cell culture to be partially synchronized to catch the cells in pro-metaphase before they condense too much. It produces 800 + bands.
Dark G bands and bright Q bands are AT rich, late replicating, gene poor, and contain LINE repeats. G light bands and dull Q bands are GC rich, early replicating, gene rich, and contain SINE/Alu sequences
Q banded Cat chromosomes Q banding FISH Telomere Cat Chromosomes
46, XX is the normal female karyotype; 46, XY is the normal male karyotype. X and Y are the sex chromosomes; all others are called autosomes. 47,XY,+21 means there is an extra #21 chromosome (a trisomy); 45,X, means there is one less chromosome, namely one sex chromosome is missing (a monosomy). Mosaicism is denoted by a "/", e.g., 45,X/46,XX indicates a Turner female with two cell lines, one normal and one missing the second X. This arises from post zygotic non disjunction, after fertilization and in mitosis of the zygote or embryo.
Haploid = n is the number of pairs of chromosomes. Most organisms are diploid, 2n, and their gametes are haploid, n. Polyploidy refers to having multiple sets of chromosomes: 3n (triploid); 4n (tetraploid). Aneuploidy refers to having (usually) one more or less of the diploid number of chromosomes: trisomies (one extra); monosomies (one less).
FISH
FISH (fluorescence in situ hybridization) has become a commonly used staining technique where known sequences of DNA are fluorescently labeled and hybridized to chromosomes.The process depends on complementary base pairing, similar to Southern blots. In situ refers to the fact that the fluorescent DNA probe is hybridized to the DNA of interphase nuclei or metaphase chromosomes which have been fixed on a slide. FISH uses locus specific DNA probes, repetitive DNA such as the centromeric alpha satellite DNA or whole chromosome painting (wcp) or spectral painting probes. FISH can be used in the identification of marker chromosomes, rings and other structural anomalies using probes which are specific for each chromosome (painting probes). Below, you can see examples of chromosome abnormalities with comparisons of GTG banding and FISH analysis. There is a picture of a 3p+ where the "+" was shown to come from chromosome 13; an iso 5q which was actually a 5 with a piece of 6; an 8 p+ where the + was a piece of 21; an 8 p+ where the + was a piece of 22. FISH is now used almost routinely for microdeletion syndromes which we will discuss later.
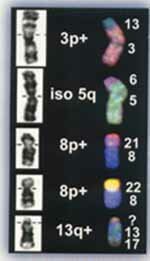
Centromeric repeat probes which bind to the unique satellite sequences give a large, diffuse but intense signal around the centromere. They can detect aneuploidies such as trisomies 21, 18, 13 even in interphase nuclei. They are not useful in detecting structural abnormalities.
Locus specific probes give a sharp, discrete signal at the relevant positions in each chromatid of a metaphase chromosome; they can be used to identify or confirm microdeletion (or macrodeletion) syndromes. An SRY probe can detect the gene responsible for testes formation even if it has been translocated to another chromosome.

Locus Specific Probes Alpha satellite centromere probes
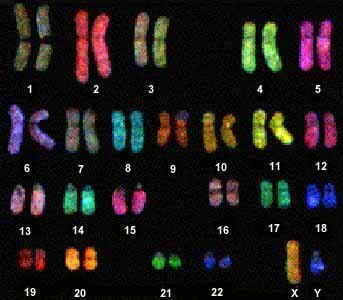
Whole Chromosome Painting (wcp)
Painting probes can uniformly decorate the entire chromosome or can be designed to paint the p or q arm, etc. The DNA sequences used are unique to the chromosome or arm painted. They can be used to identify the chromosomal origin in structural rearrangements such as translocations, rings, isochromosomes, etc. The term, "spectral karyotyping" is used to describe "whole" chromosome specific painting.
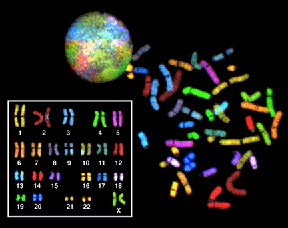

Human Chromosomes Orangutan Chromosomes

